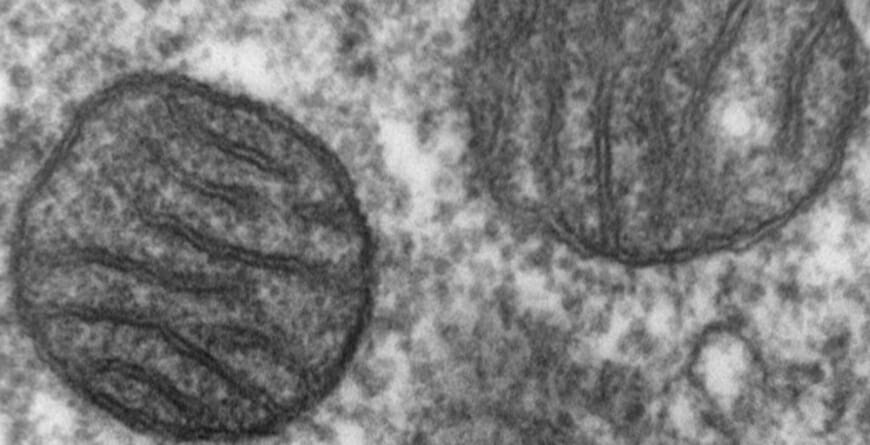
To perform myriad operations essential to life, cells require an energy source. They get it from the molecule ATP, produced through cellular respiration within specialized structures—the mitochondria.
The production of ATP is a complicated process researchers are still struggling to fully grasp. Essential to the final production of ATP are a series of five respiratory complexes, collectively known as the electron transport chain.
Scientists are working to ferret out the details of how this molecular conveyor belt works. Improved understanding could lead to new treatments for mitochondrial illnesses that result from dysfunction of the electron transport chain, including a range of serious neurodegenerative ailments. Aberrations in mitochondrial transport processes are also a source of reactive oxygen species, linked with Parkinson's disease and aging.
In a new study, Biodesign researcher Abhishek Singharoy joins lead author Chitrak Gupta to explore the first and largest of the five respiratory complexes, known as Complex I. This bulky, L-shaped complex is found within the mitochondrion, with its horizontal arm embedded in the inner mitochondrial membrane and its vertical arm protruding into the mitochondrial matrix.
Gupta and Singharoy are both researchers in the Biodesign Center for Applied Structural Discovery and ASU’s School of Molecular Sciences.
The group’s work appeared recently in the Journal of the American Chemical Society (JACS)
The purpose of Complex I is to harvest electrons from a metabolic co-factor known as NADH, and shuttle them further along the electron transport chain.
"Complex I is a result of evolution repurposing different proteins for new functions,” says Singharoy. “It's an embodiment of a redox chain and an ion pump combined together. Chitrak's simulations starting from (corresponding author Leonid) Sazanov’s experiments offer one of the closest insights on the coupling between these two universal bioenergetic paradigms. This study takes us a step closer towards the computational microscopic imaging of a complete mitochondrial membrane."
Using the latest crystal structure of Complex I, derived from the model bacterium T. thermophilus, the group was able to simulate the molecular dynamics of the complex on a microsecond scale, revealing previously elusive details of transport. To accomplish this, 17.8 μs of molecular dynamics (MD) simulations and free-energy calculations were carried out.
"We had the privilege of running our simulations on Summit, the fastest supercomputer in the world, housed at Oak Ridge National Laboratory,” Gupta says. “MD simulations of such large biological systems tend to be compute resource-intensive. Combining the power of exascale computing with sophisticated structural biology allows us to view mitochondrial respiration at the molecular level."
Joining Singharoy and Gupta were corresponding authors Christophe Chipot from the Department of Physics, University of Illinois at Urbana− Champaign and Leonid Sazanov, Institute of Science and Technology (IST), Klosterneuburg, Austria.