Using ultra-bright X-ray flashes, a team of researchers has tracked down a potential target for new drugs against sleeping sickness. The scientists have decoded the detailed spatial structure of a vital enzyme of the pathogen, the parasite Trypanosoma brucei. The result provides a possible blueprint for a drug that specifically blocks this enzyme and thus kills the parasite, as the team led by Christian Betzel from the University of Hamburg, Lars Redecke from the University of Lübeck and DESY and Henry Chapman from DESY reports in the journal Nature Communications.

Amongst this team were researchers from Arizona State University, including Petra Fromme, the director of the Biodesign Center for Applied Structural Discovery and Regents Professor in the School of Molecular Sciences.
Sleeping sickness (African trypanosomiasis) is a tropical disease caused by the parasite Trypanosoma brucei, which is transmitted by the bite of tsetse flies inhabiting much of tropical Africa. In the body, the parasite first multiplies under the skin, in the blood and in the lymphatic system and then migrates to the central nervous system. If left untreated, the disease is almost always fatal. Thanks to intensive control measures, the number of registered cases has fallen dramatically in recent years. Nevertheless, sleeping sickness is still considered one of the most significant tropical diseases. According to the World Health Organization, more than 60 million people in rural areas of sub-Saharan Africa are at risk. War, displacement and migration can cause the disease to flare up.
In the search for a possible starting point for drugs against the pathogen, the researchers had targeted a central enzyme of the unicellular organism, inosine-5'-monophosphate dehydrogenase (IMPDH). "This enzyme belongs to the central inventory of every organism and is an interesting target for drugs because it regulates the concentration of two vital nucleotides in the cell: guanosine diphosphate and guanosine triphosphate," says Redecke. "The cell needs these nucleotides to supply energy and to build larger structures such as the genome. If you interrupt this cycle, the cell dies."
The enzyme has a kind of on/off switch that is activated by the docking of the cell's own molecules. A promising approach is to block this switch with a precisely tailored molecule. In order to construct such an inhibitor, the exact spatial structure of the switch must be known. Structural biologists can determine the structure of biomolecules using X-rays. To do this, they first grow small crystals from the biomolecules, which then generate characteristic diffraction patterns when illuminated with X-rays. From these patterns the atomic structure of the crystal and its building blocks, the biomolecules, can be calculated.
This approach is often complicated by the intractability of most biomolecules against forming crystals. And if such crystals can be grown, they are usually extremely sensitive to the high-energy X-rays and are quickly destroyed. "Although the structures of numerous IMP dehydrogenases are already known, there had been no success in growing crystals of the Trypanosoma brucei version of the enzyme", reports Betzel, who is also a researcher in the Cluster of Excellence CUI: Advanced Imaging of Matter at the University of Hamburg and DESY.
The team therefore chose an alternative route: the group of co-author Michael Duszenko at the University of Tübingen induced certain insect cells to crystallize biomolecules within them. Using this so-called in cellulo crystallization, the same team had already deciphered another key enzyme of the sleeping sickness pathogen, cathepsin B, which is also a potential drug target. It turned out that the altered insect cells also produce crystals of the dehydrogenase now investigated. These crystals form tiny needles around 5 thousandths of a millimeter (5 micrometers) thick and up to 70 micrometers long, so that they protruded from the producing cells.
The in-cellulo crystals are so small that very bright X-rays are required to analyze them. The larger a crystal is, the more atoms within it can scatter X-rays, causing a better diffraction pattern. The researchers therefore used the LCLS X-ray laser at the SLAC National Accelerator Laboratory in the US for the analysis. "X-ray lasers generate extremely intense flashes," explains Chapman, who is a Lead Scientist at DESY in the Center for Free-Electron Laser Science CFEL and one of the spokespersons of the Cluster of Excellence CUI: Advanced Imaging of Matter. "Although the sensitive crystals evaporate immediately, they first generate a diffraction pattern from which the structure can be obtained." The method used here to exploit these properties, called serial femtosecond crystallography, was developed earlier by many of the researchers involved in this study and named one of the top ten breakthroughs of the year by Science magazine in 2013.
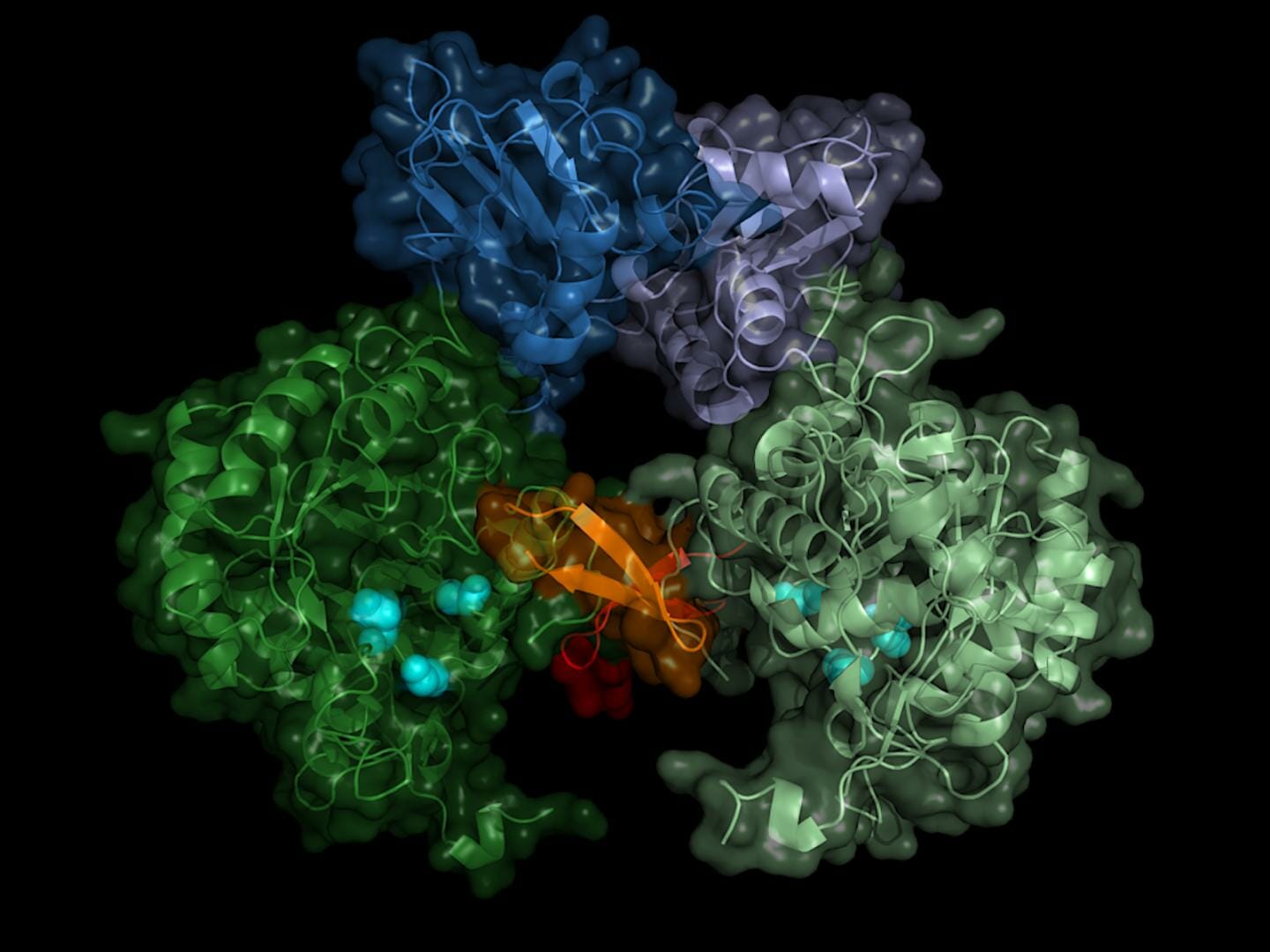
The team recorded the diffraction patterns of more than 22,000 microcrystals and was able to calculate the spatial structure of the enzyme with an accuracy of 0.28 millionths of a millimeter (nanometer) - this corresponds roughly to the diameter of an aluminum atom. "The result does not only show the exact structure of the enzyme switch, the Bateman region, but also which molecules of the cell activate the switch and how these so-called co-factors bind to the enzyme switch," reports Karol Nass who performed this work during his PhD studies at DESY. He now works at the Paul Scherrer Institute in Switzerland and is, together with Redecke, a main author of the publication.
The switch is operated by the molecules adenosine triphosphate (ATP) and guanosine monophosphate (GMP). "The advantage of our method is not only that we are able to investigate the enzyme at room temperature, at which the enzyme naturally operates, but also that during in cellulo crystallization the natural co-factors bind to the enzyme," said Betzel. According to the researcher, the data might now provide an approach for inhibiting the parasite's IMP dehydrogenase. "One could think of constructing a kind of clasp that would cover the binding sites of both co-factors, for example."
However, a remaining challenge is to design the IMP dehydrogenase inhibitor in such a specific way that it blocks the parasite's enzyme, but not the human enzyme. If this is successful, the method could potentially be extended to other pathogens, explains Betzel. "Other parasites have a very similar structure, and it might also be possible to attack those via the respective IMP dehydrogenase. The enzyme is a very interesting target for drugs, for example against the fox tapeworm or the elephantiasis pathogen."
The universities of Hamburg, Lübeck and Tübingen, the Russian Academy of Sciences, Arizona State University, the Lawrence Livermore National Laboratory in the USA, the Max Planck Institute for Medical Research, the US National Accelerator Laboratory SLAC, the University of Gothenburg and DESY were involved in this research.
###
Reference:
In cellulo crystallization of Trypanosoma brucei IMP dehydrogenase enables the identification of genuine co-factors; Karol Nass, Lars Redecke et al.; Nature Communications, 2020; DOI: 10.1038/s41467-020-14484-w